Demonstration and testing of checkpointing and restarting openCARP
See code in GitLab.
Author: Gernot Plank gernot.plank@medunigraz.at and Anton J Prassl anton.prassl@medunigraz.at
Efficient organization of in silico experiments by checkpointing
Like wet-lab in vivo or in vitro experiments, simulated in silico experiments implement protocols where the system's response to a set of perturbations is interrogated. The perturbation of the reference state, typically a system limit cycle under a given condition such as pacing at a constant cycle length or a pacemaker driving activation at a constant rate, is variable, governed by a set of experimental parameters. For instance, an S2 stimulus is delivered to test whether reentry can be induced, but the success or failure of induction is dependent on the coupling interval, the strength and duration of the stimulus and the stimulus location. This spans a parameter space that must be sampled in an experiment to determine under which conditions an S2 stimulus is able to induce reentry. How is such an experimental protocol efficiently implemented in an in silico experiment?
Unlike wet-lab experiments, in silico experimentation offers potential for protocol optimization. Wet-lab experiments are linear in time, every protocol must be implemented from start to end, necessitating the repetition of all segments of the experiment. In in silico experiments a protocol can be broken down in segments where the overlapping segments do not have to be repeated. Implementation of segmented execution of experimental protocols requires the simulator to be able to mark the end of a segment, and to continue a simulation from this mark. This capability is referred to as checkpointing and restarting. openCARP offers a broad spectrum of checkpointing techniques that can help, if wisely used, to reduce compute costs substantially.
As an example to showcase the application of checkpointing, we consider an S1-S2 protocol for testing reentry inducibility. We consider a tissue sheet where we pace the left hand side of the sheet at a given pacing cycle length to a limit cycle (using ForCEPSS, see this example for details). We deliver an S2 stimulus then that covers the left upper hand quarter of the tissue. Using a sufficiently strong stimulus strength, the stimulus will activate the tissue in excitable regions but will fail in regions not fully recovered. At the intersection between critical stimulus strength line with the critical recovery isoline, a phase singularity will emerge around which a rotor forms driving a functional reentry. In this simple experiment, the only experimental variable we will explore is the coupling interval of the S2 stimulus. That is, we seek to find the CIs which induce by evaluating the function
where we interrogate a range of CI by a chosen sampling: CI = [150, 160, 170, …, 240, 250].

In a wet lab experiment, this requires the execution of all 7 protocol variations over the entire duration of the protocol. That is, after inducing or not inducing an arrhythmia the tissue must be reset (if induction succeeded the arrhythmia must be terminated first), the entire limit cycle pacing must be repeated, before the next CI can be applied.
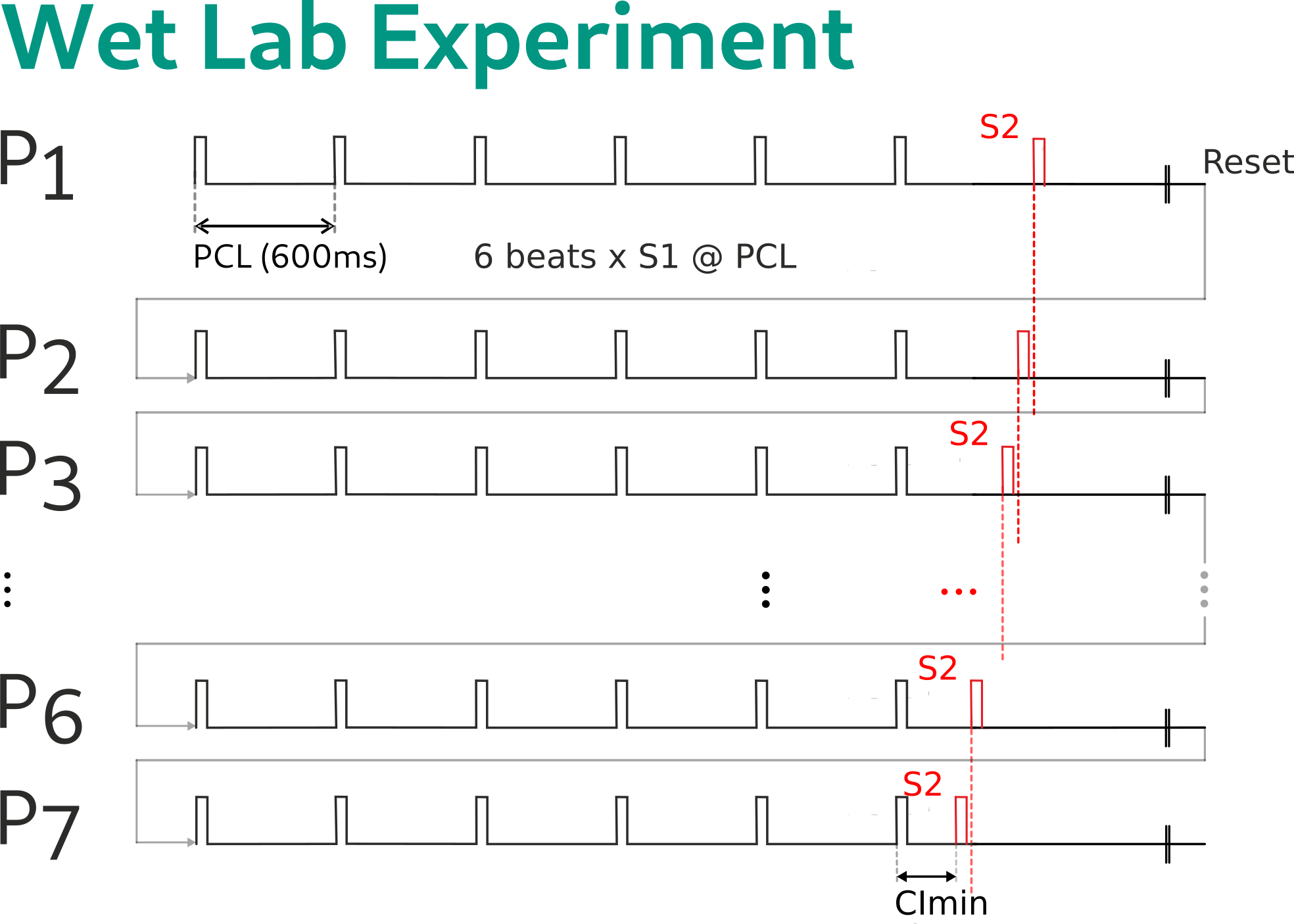
This is fundamentally different in an in silico experiment. Under this protocol, we identify the segments that are identical for each protocol. In the simplest case, we identify the S1 phase as a joint segment that is identical for all 7 experiments. As such, this part of the computation is not repeated. For instance, we delivered 6 S1 stimuli at a pacing cycle length of 600 ms, we checkpoint therefore the end of limit cycle pacing at t=3100ms, and restart the remaining part of the experimental protocol from there.
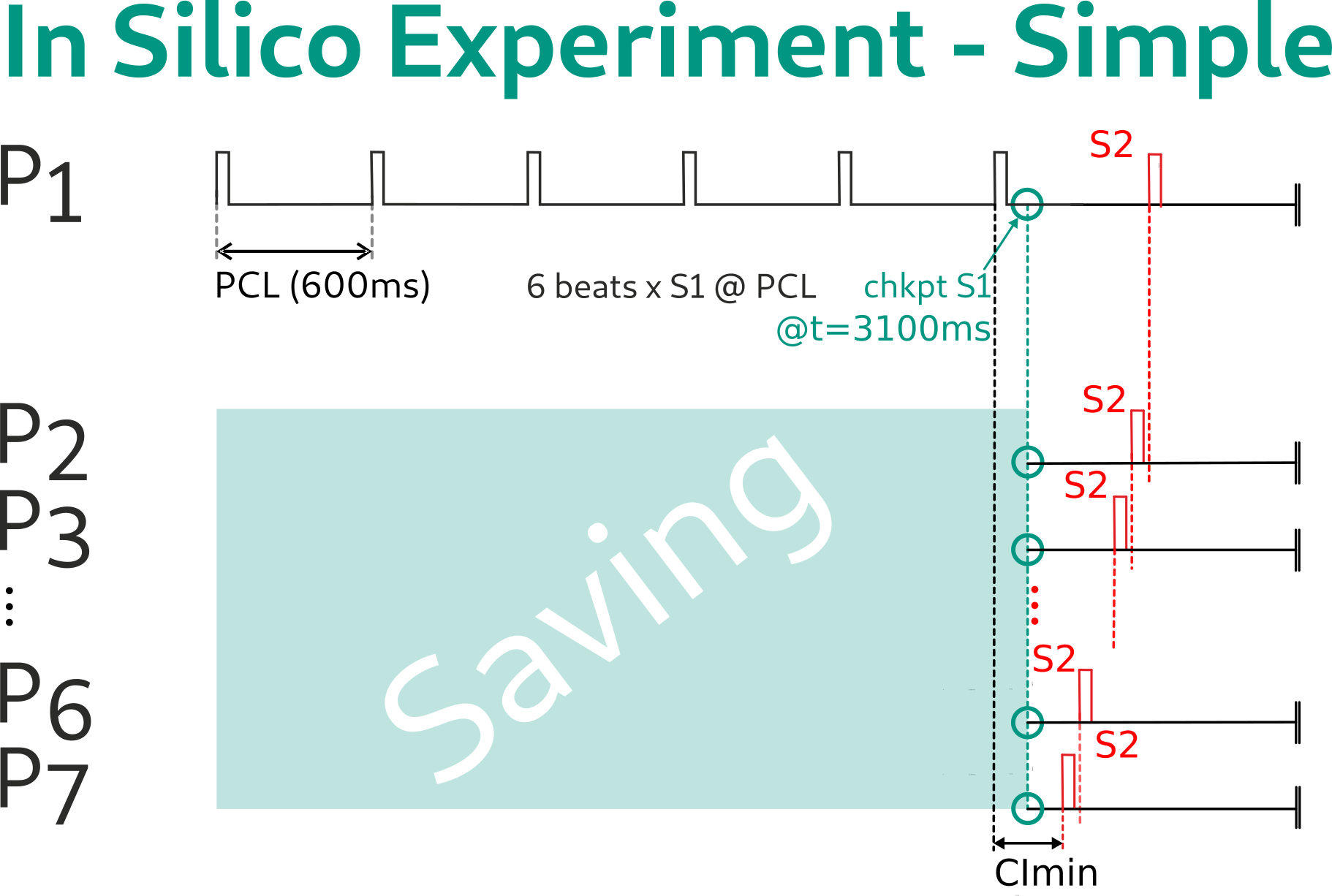
The computational savings are then the simulation of 6 x 3100 ms. For a larger scale setup, such as whole organ simulations, the savings may be days of core hours on a supercomputer. Such a checkpoint is executed in openCARP as follows:
-num_tsav 1
-tsav[0] 3100With these parameters, openCARP will write a checkpoint into the output directory specified by -simID named state.3100.0.roe, which can be used for restarting the simulation from there. This is done by adding the following command to the par file:
-start_statef /path_to_simID/state.3100.0An even more efficient approach is to run the simulation beyond the limit cycle period and simulate parts of the S2 cycle, where we checkpoint all instants in time at which the S2 will be delivered. This is implemented by checkpointing at the instants of the end of the limit cycle period + the CI of the S2. This is achieved by:
-num_tsav 11
-tsav[0] 3150
-tsav[1] 3160
-tsav[2] 3170
-tsav[3] 3180
-tsav[4] 3190
-tsav[5] 3200
-tsav[6] 3210
-tsav[7] 3220
-tsav[8] 3230
-tsav[9] 3240
-tsav[10] 3250As the intervals between checkpoints are constant, a more concise definition can be used:
-chkpt_start 3150
-chkpt_intv 10
-chkpt_stop 3250Both approaches yield the same results with the only difference being the naming of the checkpoint. Checkpoints generated with the tsav mechanisms are named state*, those written with the checkpoint interval specification are named checkpoint*. Both mechanisms can be arbitrarily mixed together, including dumping of checkpoints at the same time, the mechanisms are entirely independent.
Using the latter approach, the simulation can be restarted exactly at the time of the S2 delivery, yielding even more savings:
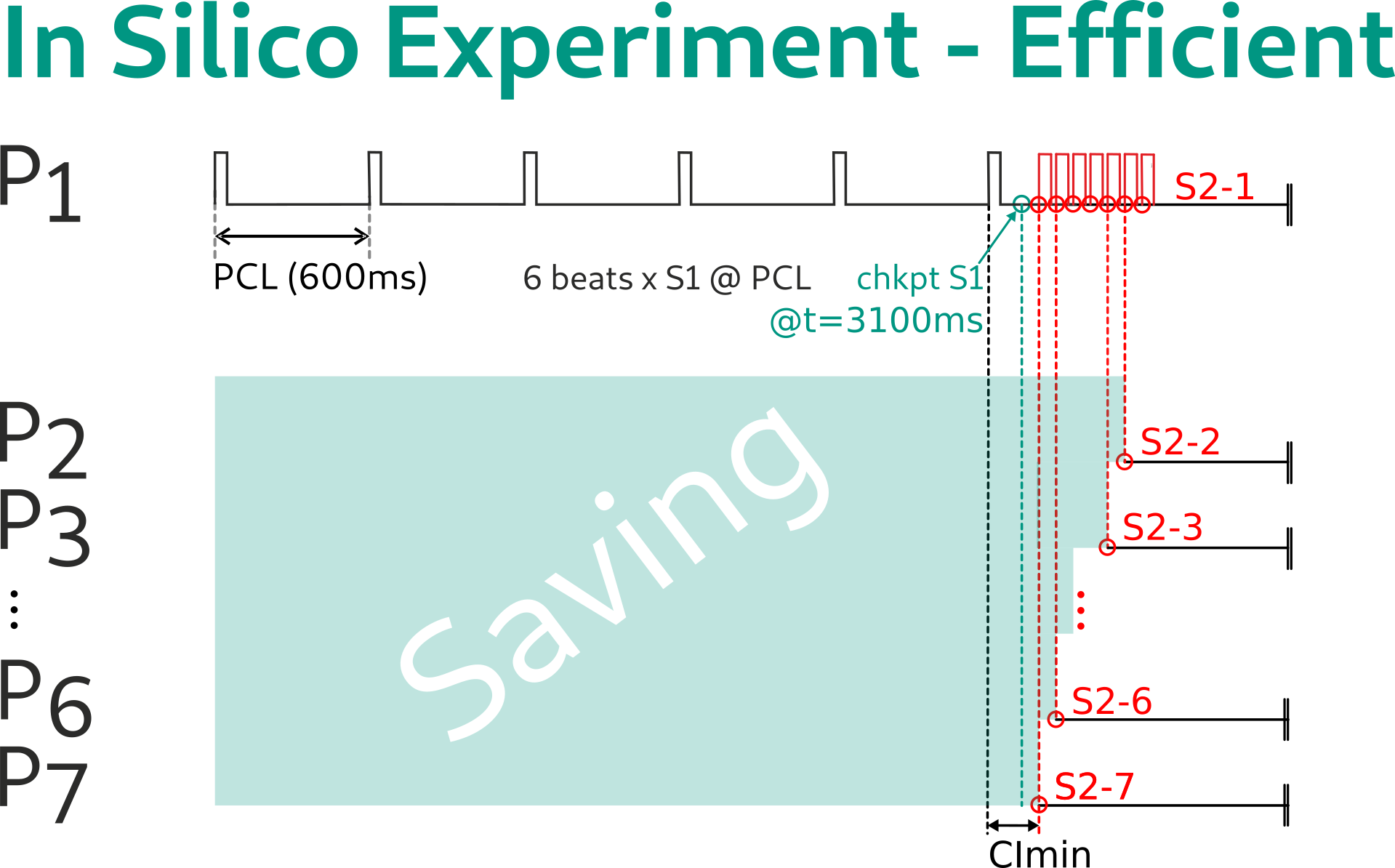
Now the outcome of all CIs can be efficiently determined minimizing redundancy in computations. Evaluating the outcome of the simulation requires the construction of the function \(p(CI)\).
There are three possible outcomes:
- The S2 is premature and does not capture, no new wavefront is induced.
2) The S2 is too late, the tissue around the S2 is fully recovered, thus the S2 induces an activation, but no reentry is observed. 3) The S2 intersects with the recovery isoline and induces a rotor.
For scenario 3), we would like to measure also maintenance of the rotor. We are only interested in stable rotors that are maintained over a prolonged observation period of 2000 ms, transient rotors that die out earlier than this should be ignored. A straightforward implementation of this protocol requires the simulation of 2000ms in all 7 cases, independently of the scenario 1), 2) or 3).
An efficient way of evaluating the outcome is the use of a sentinel mechanisms. In our simulation, we would like to identify whether new activation events (action potentials) occur in our tissue. If there are no new activation events detected over a given observation period, we terminate the simulation and write a checkpoint (thus keeping the option to continue the simulation if needed). We specify a sentinel by
-sentinel_ID 0
-t_sentinel_start 3100
-t_sentinel 40Telling the simulator to start tracking activation events at time t=3100 and terminate the simulation if there are no new activations over a period of 40 ms. In scenarios 1) and 2), simulations will terminate after 40 ms as there are no new activation wavefronts emerging. In scenario 3), simulations will terminate 40 ms after the rotor dies out, or at the end of the protocol, if the rotor is maintained. As an example, we showcase these two outcomes:
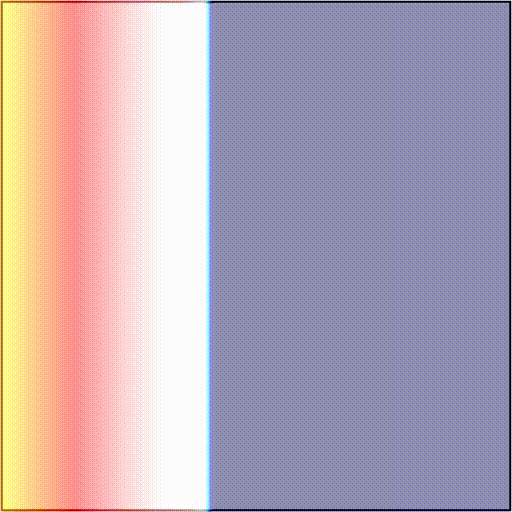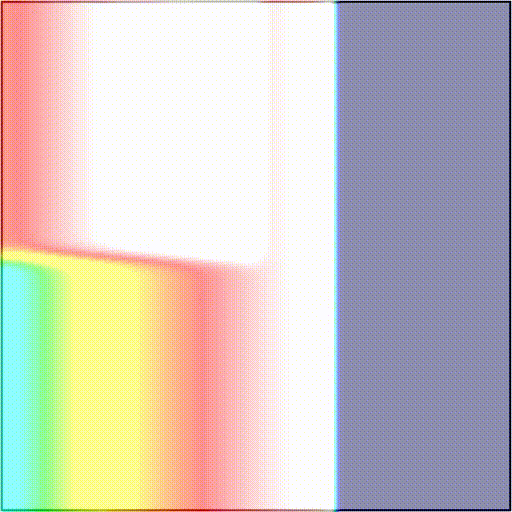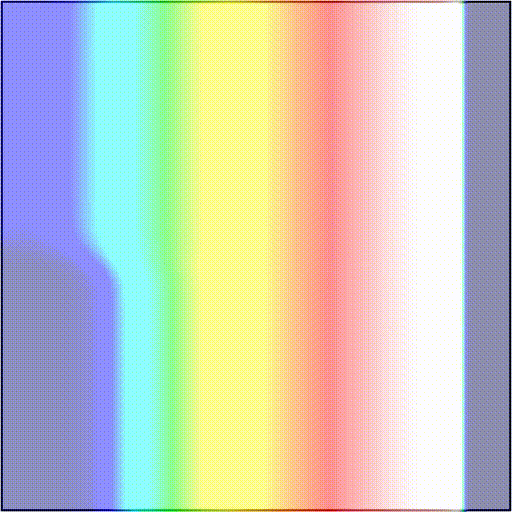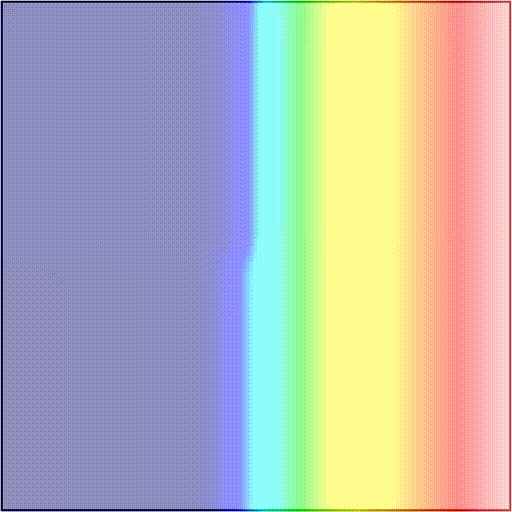
The presence and timing of the sentinel checkpoint can be interrogated after a simulation to determine the outcome of the protocol.
Recent questions tagged carputils, sentinel, checkpoint, statefile, forcepps
There are tagged with carputils, sentinel, checkpoint, statefile, forcepps.
Here we display the 5 most recent questions. You can click on each tag to show all questions for this tag.
You can also ask a new question.
